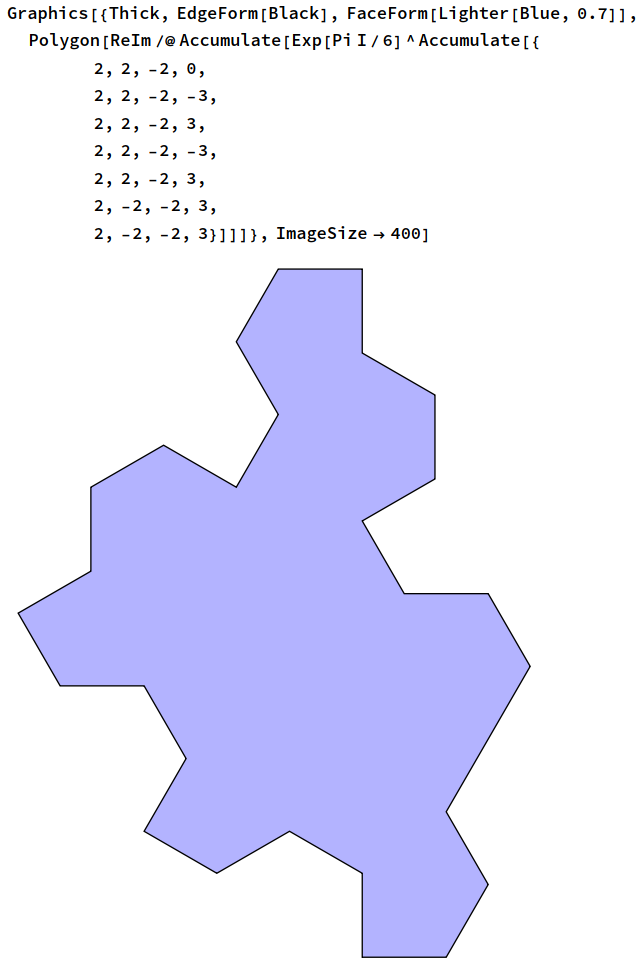
I’ve been spending some of my time recently developing a tool called FPSan in collaboration with Pawel Szczerbuk. It’s implemented as a Triton compiler pass, but has none of the desirable properties expected of a compiler pass: in particular, it doesn’t preserve functionality, it makes things slower, and it’s hitherto completely undocumented. (On the latter point, Pawel has an open PR adding documentation.)
Its purpose is to make it easier to verify algebraic equivalence of programs written in Triton that involve floating-point arithmetic. The key problem is that, in floating-point arithmetic, algebraic laws such as associativity do not hold exactly: in general, (a + b) + c need not equal a + (b + c). As such, if you rewrite a program to take advantage of this, e.g. to replace a sequential summation loop with a parallel tree-shaped reduction, the program will no longer behave completely identically.
FPSan can be viewed as an idempotent function on the space of programs that replaces all floating-point operations with (completely different!) integer operations, such that if f and g are algebraically equivalent programs then FPSan(f) and FPSan(g) produce identical results when given identical inputs.
More formally, conditional on the real version of Schanuel’s conjecture, this holds provided that the programs f and g have the following properties:
- each program implements an arithmetic circuit on its floating-point inputs, and the control flow is independent of those floating-point inputs;
- the arithmetic circuit only consists of inputs, outputs, the constants {-1.0, 0.0, +1.0}, the ring operations {−, +, ×}, and the exponential function exp.
These operations may seem somewhat restrictive, but it already encompasses a vast range of the more common GPU kernels involved in machine learning: matrix multiplications and [the bulk of] self-attention are covered by FPSan’s guarantees.
The proof is deferred to the end of this article to avoid derailing the discussion. This is quite possibly the only compiler sanitiser whose correctness depends on an extremely difficult unsolved problem in transcendental number theory.
Implementation
Specifically, FPSan constructs a bijective ’embedding function’ φ from the set of IEEE-754 single-precision floats (there are 2^32 of them) to the ring of integers modulo 2^32. The function φ is implemented as follows:
- encode the float as a 32-bit word using the IEEE-754 encoding;
- the uppermost bit (sign bit) is preserved;
- for the remaining 31 bits, we apply a mod-2^31 multiplication by an odd constant, then a xorshift, then another mod-2^31 multiplication by an odd constant, and finally (if the sign bit was set) take the two’s complement;
- interpret the 32-bit word as an integer modulo 2^32.
It’s designed to mix the bits reasonably well whilst having the properties that φ(−x) = −φ(x) for all nonzero x, φ(0.0) = 0, and φ(1.0) = 1. The ‘negative zero’ float gets mapped to 2^31, which is the other additively self-inverse element of the ring of integers modulo 2^32.
With this function, FPSan replaces:
- floating-point addition fadd(x, y) with φ^-1(φ(x) + φ(y));
- floating-point subtraction fsub(x, y) with φ^-1(φ(x) − φ(y));
- floating-point multiplication fmul(x, y) with φ^-1(φ(x) × φ(y));
- floating-point exponentiation exp(x) with φ^-1(C^φ(x)) where C is a particular constant that’s congruent to 5 (mod 8).
The last of these definitions makes use of the structure of the multiplicative group of integers modulo 2^32. In particular:
- only the 2^31 odd elements, those that are 1 (mod 2), are invertible and therefore belong to the multiplicative group;
- of those, the 2^30 elements that are 1 (mod 4) form a cyclic group under multiplication;
- of those, the 2^29 elements that are 5 (mod 8) are generators, or equivalently have maximal period, which is why we choose C to be 5 (mod 8).
The map x → C^x is well-defined mod 2^32, because C^(2^32) = 1 (mod 2^32), so C^x (mod 2^32) depends only on the value of x mod 2^32. This works only because our modulus is a power of two; for an arbitrary modulus n, the exponent of the multiplicative group does not in general divide n.
The rewritten versions of fadd, fsub, fmul, and exp evidently obey all of the ring axioms, the identity exp(fadd(x, y)) = fmul(exp(x), exp(y)), and the relation exp(0.0) = 1.0.
Mixed-precision functionality
FPSan constructs an analogue of the embedding function φ for arbitrary floating-point datatypes, mapping into an integer ring of the same cardinality. To downcast from j-bit to k-bit precision, we embed our high-precision j-bit float into the ring of integers mod 2^j, then take the image mod 2^k, then unembed as a low-precision k-bit float. Upcasting is the reverse, where we choose the “sign-extending” lift from the integers mod 2^k to the integers mod 2^j; this in particular means that the constants {-1, 0, 1} survive arbitrary casting between different precisions. An upcast followed by a downcast induces the identity map; the reverse is not true because downcasting necessarily destroys information.
Constructing the multipliers in the embedding function and its inverse requires being able to compute efficient inverses modulo 2^k; we do this using ceil(log2(k)) iterations of the 2-adic Newton’s method.
Pawel wrote the rules for converting Triton’s mixed-precision matrix multiplication primitive, tl.dot, into the FPSan equivalent by expanding it into floating-point scalar operations. The mixing functions φ and φ^-1 only need to be applied to each input and output element, with the core of the matrix multiplication only involving int32 multiplication and addition.
The proof
Now for the fun part: the proof that Schanuel implies the desired properties of FPSan.
Suppose that we have two arithmetic circuits f and g over the reals, each consisting only of inputs, outputs, the constants {-1, 0, +1}, the ring operations {−, +, ×}, and the exponential function exp. Suppose moreover that they’re equivalent in that they implement the same function from to
.
Assuming the real version of Schanuel’s conjecture, we have that the subring X of R generated by {0, −, +, ×, exp} is isomorphic to the free exponential ring on no generators, as proved in Macintyre 1991. If the circuits are equivalent over R, then they’re necessarily equivalent when we restrict to X, and by the isomorphism that means that they’re equivalent in the free exponential ring on no generators.
The ring of integers mod 2^32 together with the unary function C^x (where C is a particular constant that’s congruent to 5 (mod 8)) is a quotient of the free exponential ring on no generators; in particular, we can construct a surjective homomorphism θ from the free exponential ring on no generators to the ring of integers mod 2^32 by setting θ(exp(x)) = C^θ(x).
It follows, therefore, that the circuits remain equivalent under FPSan, as that just endows the floats with the structure of an exponential ring by pulling back the operations through the embedding function φ.
Sine and cosine
We also implement analogues of sine and cosine over the 2-adic integers by taking the real and imaginary parts of (−3/5 + 4/5 i)^n in the quadratic extension obtained by adjoining a formal symbol i satisfying i^2 = −1. These satisfy the trigonometric angle sum and difference identities, along with the usual norm identity sin(x)^2 + cos(x)^2 = 1.
The result (that any valid algebraic identity involving {0, 1, −, +, ×, exp} that holds over the reals also holds under FPSan, assuming Schanuel’s conjecture) can be strengthened: it remains true when sin and cos are included.
We’ll define the following two sequences of rings:
- an ascending chain of subrings of the reals, {Y_0, Y_1, Y_2, …}, where Y_0 = Z and Y_{n+1} is the ring generated by adjoining Y_n with exp(x), sin(x), and cos(x) for all x in Y_n;
- an ascending chain of subrings of the complex numbers, {W_0, W_1, W_2, …}, where W_0 = Z[i] and W_{n+1} is the ring generated by adjoining W_n with exp(x) for all x in W_n.
We can show by induction on n that W_n is exactly Y_n[i] (and in particular Y_n is the real part of W_n). In particular, assuming this holds for n−1, we have:
- if x is in Y_{n−1}, then exp(x), cos(x) = (exp(ix) + exp(-ix))/2, and sin(x) = (exp(ix) – exp(-ix))/(2i) are all clearly in W_n;
- if z is in W_{n−1}, so its real and imaginary parts a and b are in Y_{n−1}, then the real and imaginary parts of exp(z) are exp(a) cos(b) and exp(a) sin(b) which reside in Y_n.
Defining Y to be the union of Y_n for all n, and similarly defining W to be the union of W_n for all n, we have W = Y[i].
We can now repeat Macintyre’s proof idea but on the sequence of rings W_n: assuming the complex Schanuel’s conjecture, W is the free exponential ring generated by an i satisfying i^2 = −1. Any algebraic relation in Y involving {0, 1, −, +, ×, exp, cos, sin} can be converted to an algebraic relation in W involving {0, 1, i, −, +, ×, exp}, and must hold in any exponential ring where i satisfies i^2 = −1. The result follows.